ANALYTICAL CHEMISTRY LABORATORY
of Molecular Phytopathology and Mycotoxin Research Group

|

|

|

|
- Mycotoxins
- Plant metabolites
- Phytohormones
- Pesticides
- Metabolic profiles
- Purification of standards
- Volatile compounds
What we analyze
- HPLC-Q-TOF and HPLC-QQQ
- HPLC-IT 500-MS (retired)
- HPLC-QQQ L1200 (retired)
- GC-FAIMS (retired)
- Flash chromatography
- Preparative TLC
- Preparative HPLC
- HPLC-FD/DAD
- Capillary electrophoresis (retired)
- Spectrofluorometry
Our instruments
Our chemical analysis: a synopsis
We use chemical analysis in our research and provide analytical services to our partners and collaborators. We mainly use HPLC coupled with optical (DAD, FD, ELSD) and mass spectrometric (ion trap, Q-TOF and triple quadrupole) detectors. Flash chromatography, preparative HPLC and Clevenger apparatus are used for the purification of fungal and plant metabolites and for standards that we synthesize in the lab. Further equipment include rotary evaporators, vacuum concentrators, a portable GC-FAIMS, a spectrofluorometer, a microplate fluorometer and different electrophoreses (PAGE, DGGE, CE).
Contact
Dr. Mohammad Alhusseinphone: (0)551-23471email: "mohammad dot alhussein at uni dash goettingen dot de"
Techniques
Mycotoxin analysis
HPLC-MS/MS in MRM mode is used for mycotoxin analysis. Because we work with diverse matrices such as grains, green plants tissue and insects, we use matrix-matched standards and isotopically labelled internal standards. In the past, we worked with a triple quadrupole 1200L from Varian and an ion trap (500-MS), excellent and reliable machines that served us for over a decade. A typical results is this example from 2005. Since 2018, we are using a new system equipped with a triple quadrupol from Agilent.
Since 2018: new systems for targeted and non-targeted HPLC-MS analysis
We acquired two new MS detectors from Agilent: A time-of-flight (TOF) detector for non-targeted analysis and a triple quadrupole for quantitative analysis of phytohormones, mycotoxins and other metabolites. An our ion trap from Varian provides MS^3 and higher-order fragmentation.
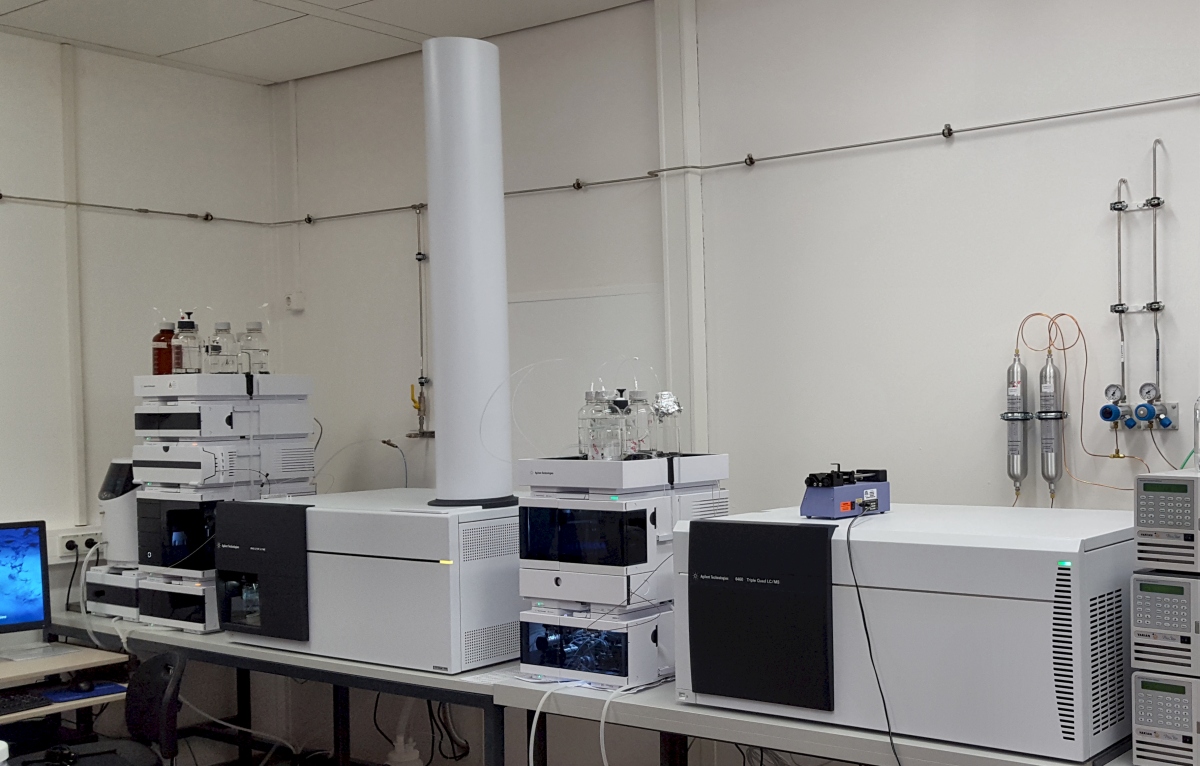

New MS systems: HPLC-Q-TOF (6545, Agilent) and HPLC-QQQ (triple quadrupol 6460, Agilent)
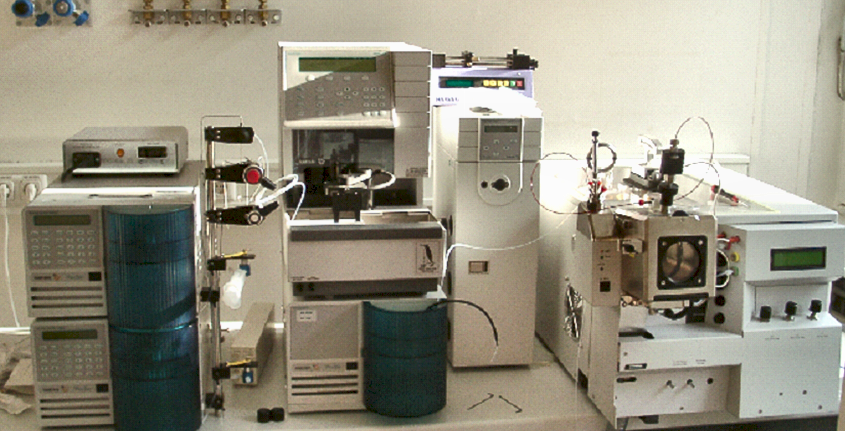

Our first MS was triple quadrupole LC1200 from Varian, which served us from 2004 till 2015.


On the left you see our second MS, which is ion trap MS-500 from Varian, with an open housing. We purchased it in 2006 and still use it (November 2018). On the right you see triple quadrupole LC1200 with an open housing.
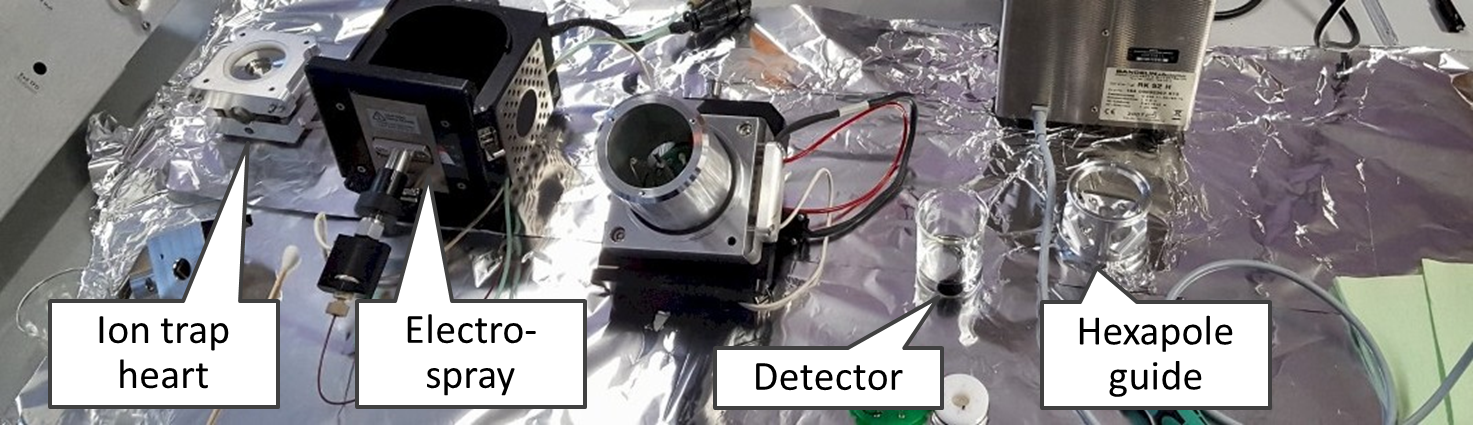

This is an ion trap disassembled into the smallest parts (users are not expected to do this!).
Although HPLC-MS/MS in MRM is a golden standard for multi-mycotoxin analysis today, in certain situations HPLC-FD is preferable.
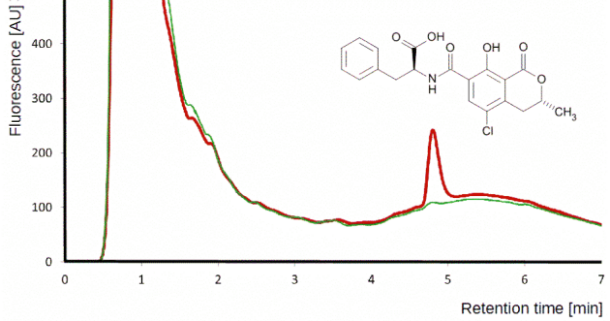
Detection of ochratoxin A by HPLC-FD
HPLC-DAD proved useful in some mycotoxin projects, too, for instance in Dr. Rosine Suchfort's work on enniatins. Another application of HPLC-DAD is Rasoul Abousaeedi's research on fusaric acid:
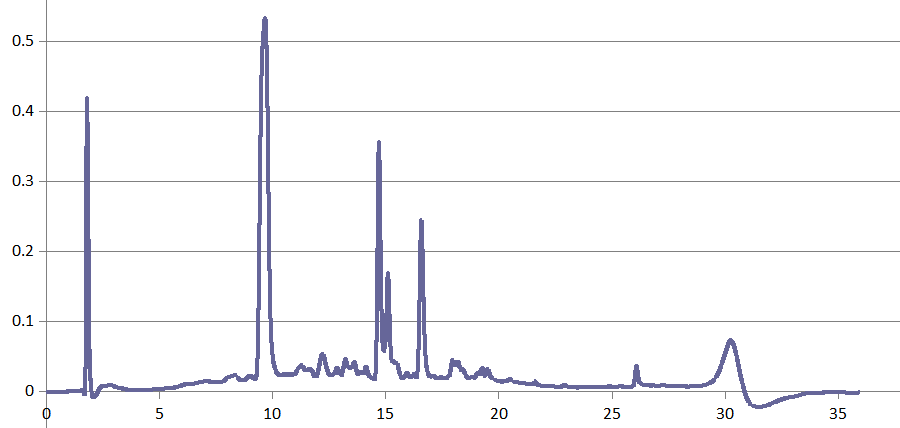
Transformation products of fusaric acid resolved by HPLC-DAD (R. Abousaeedi)
Examples of publications relying on our mycotoxin analysis by HPLC-MS/MS:
ten Bosch L, Pfohl K, Avramidis G, Wieneke S, Viöl W, Karlovsky P: Plasma-based degradation of mycotoxins produced by Fusarium, Aspergillus and Alternaria. Toxins (2017) 9/97 (12 pages) OPEN ACCESS
Guo Z, Pfohl K, Karlovsky P, Dehne HW, Altincicek B (2016) Fumonisin B1 and beauvericin accumulation in wheat kernels after seed-borne infection with Fusarium proliferatum. Agric Food Sci 25: 138-145. OPEN ACCESS
Amato B, Pfohl K, Tonti S, Nipoti P, Dastjerdi R, Pisi A, Karlovsky P and Prodi A (2015) Fusarium proliferatum and fumonisin B1 co-occur with Fusarium species causing Fusarium Head Blight in durum wheat in Italy. Journal of Applied Botany and Food Quality 88: 228-233. OPEN ACCESS
Plant secondary metabolites
HPLC-DAD is well suitable for most plant metabolites. With the help of pure standards every metabolite absorbing UV light can easily be quantified. HPLC-DAD is however useful for nontargeted analysis, too. UV absorption spectra combined with relative retention time are offten sufficient to point at the chemical identity of such metabolite even when standards are not available, as shown on the following figure. Dr. Franz Hadacek maintains a library of UV spectra:
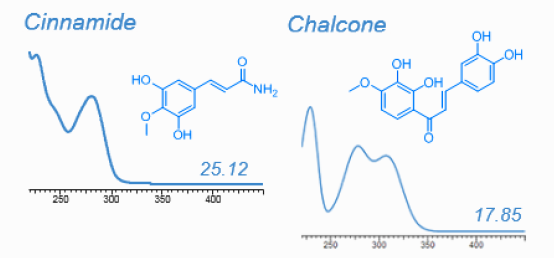
Metabolites identified in root exudates of Arabidopsis thaliana (Dr. Pervin Akter and Dr. Franz Hadacek)
HPLC-DAD can often reveal differences among complex mixtures:
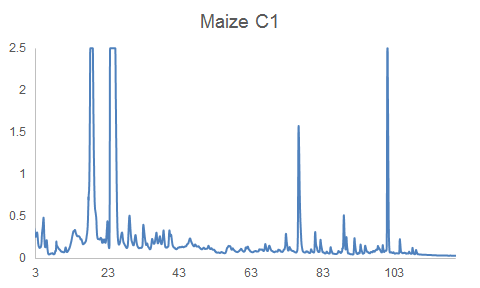
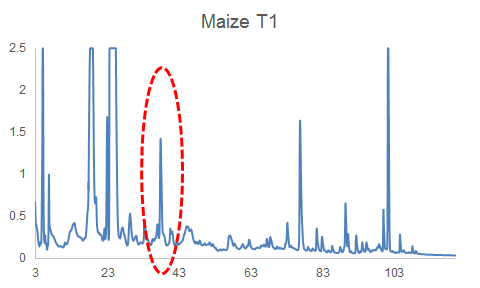
HPLC-DAD chromatograms of root exudates of maize under normal conditions (left) and under stress (right) (Dr. Pervin Akter and Dr. Franz Hadacek)
Phytohormone analysis
We use a modified ether extraction protocol with internal isotopically labelled standards, followed by HPLC-MS/MS. Examples:
Wietzke A, Westphal C, Gras P, Kraft M, Pfohl K, Karlovsky P, Pawelzik E, Tscharntke T, Smit I: Insect pollination as a key factor for strawberry physiology and marketable fruit quality. Agriculture, Ecosystems and Environment (2018) 258: 197-204.
Ulferts S, Delventhal R, Splivallo R, Karlovsky P, Schaffrath U (2015) Abscisic acid negatively interferes with basal defence of barley against Magnaporthe oryzae. BMC Plant Biology 15/7.OPEN ACCESS
Häffner E, Karlovsky P, Splivallo R, Traczewska A, Diederichsen E (2014) ERECTA, salicylic acid, abscisic acid and jasmonic acid modulate quantitative disease resistance of Arabidopsis thaliana to Verticillium longisporum. BMC Plant Biology 14/85. OPEN ACCESS
Ratzinger A, Riediger N, von Tiedemann A., Karlovsky P (2009) Salicylic acid and salicylic acid glucoside in xylem sap of Brassica napus infected with Verticillium longisporum. J Plant Res 122:571-579. OPEN ACCESS.
Pesticide analysis
So far our lab provided pesticide analysis (fungicide residues and degradation products of glyphosate) for three divisions of the Faculty of Agriculture and one division of the Faculty of Forest Sciences and Forest Ecology. The demand on pesticide analysis is likely to increase.
Metabolic profiling: nontargeted comparative analysis of metabolic profiles
Plant extracts, fungal culture supernatants, root exudates etc. are separated by HPLC on a reverse phase column, ionized by electrospray and detected by a time-of-flight (TOF) MS detector. Our data processing pipeline relies on commercial software, Perl scripts and open-source plattforms such as XCMS.
If the metabolite has already been known, its identity can often be confirmed by high-resolution m/z of a molecular ion and its fragments, sometimes supported by a UV absoption spectrum. Dr. Franz Hadacek maintains a database of UV spectra of metabolites that we have studied, which allows him to identify these metabolite when we encounter them again based solely on their retention time and UV spectrum. Below are examples of his analysis of maize root exudates:
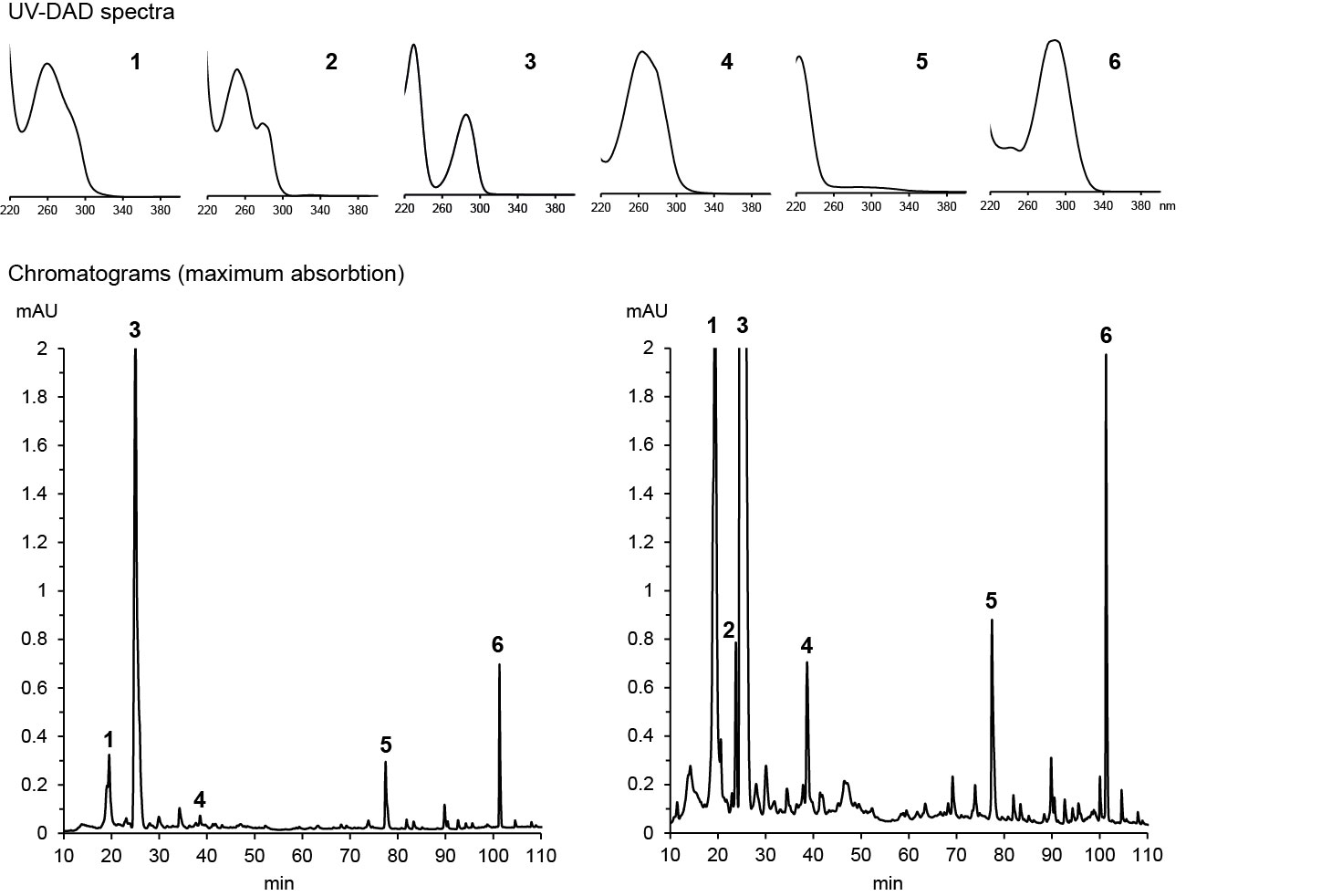
HPLC-DAD chromatograms of root exudates of maize under normal conditions (left) and under stress (right) with UV spectra recorded at major peaks (Dr. Pervin Akter and Dr. Franz Hadacek)
Information about biosynthetic capacity of the strain/species used as a source is helpful. Our Perl scripts search available databases automatically but unfortunately most metabolites relevant for the projects that we have been supporting are underrepresented in databases.
Often purification is necessary for the identification of compounds responding to a treatment (see below). Because the efficiency of ionization and thus intensity of the MS signal varies widely with structure of the analyte, wrong targets for purification were often selected in the past, resulting in purified metabolites in amounts insufficient for structure elucidation and biological assays. The evaporative light scattering detector (ELSD) will solve this problem.
Examples of papers in which we used nontargeted metabolic profiling:
Chatterjee S, Kuang Y, Splivallo R, Chatterjee P, Karlovsky P (2016) Interactions among filamentous fungi Aspergillus niger, Fusarium verticillioides and Clonostachys rosea: fungal biomass, diversity of secreted metabolites and fumonisin production. BMC Microbiology 16/83 (13 pp).OPEN ACCESS
Döll K, Chatterjee S, Scheu S, Karlovsky P, Rohlfs M (2013) Fungal metabolic plasticity and sexual development mediate induced resistance to arthropod fungivory. Proceedings of the Royal Society B 280:20131219. FREE ACCESS
Khorassani R, Hettwer Z, Ratzinger R, Steingrobe S, Karlovsky P, Claassen N (2011) Citramalic acid and salicylic acid in sugar beet root exudates solubilize soil phosphorus. BMC Plant Biology 11/111. OPEN ACCESS.
Purification and synthesis of standards
For many metabolites commercial standards for HPLC analysis are not available. We purify such standards from plant and fungal extracts using flash chromatography with gradient elution, preparative HPLC, Clevenger apparatus and preparative TLC. Typically flash chromatography on silica gel with cyclohexane/ethyl acetate/methanol gradient elution is the first step, followed by Sephadex LH20 and/or preparative HPLC. Since Nahid (Anna) Najafi joined our lab, we started synthesizing certain standards, for instance 13C-labeled glucoside of salicylic acid and cyclobrassinin.
Apart from the preparation of standards, purified and/or synthetic metabolites are needed for studies of toxicity and enzymatic transformations. Below you see purification of enniatins as an example:
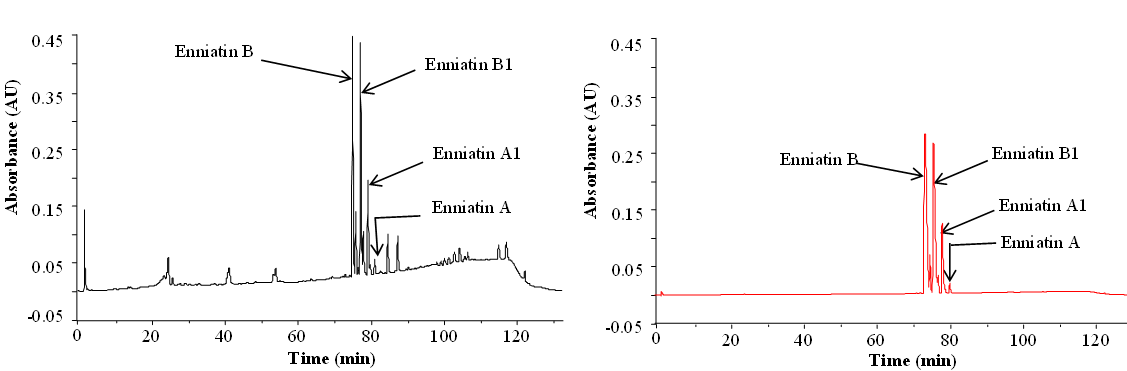
Enniatins from Fusarium tricinctum as a raw extract (left) and after purification (right) (Dr. Rosine Suchfort)
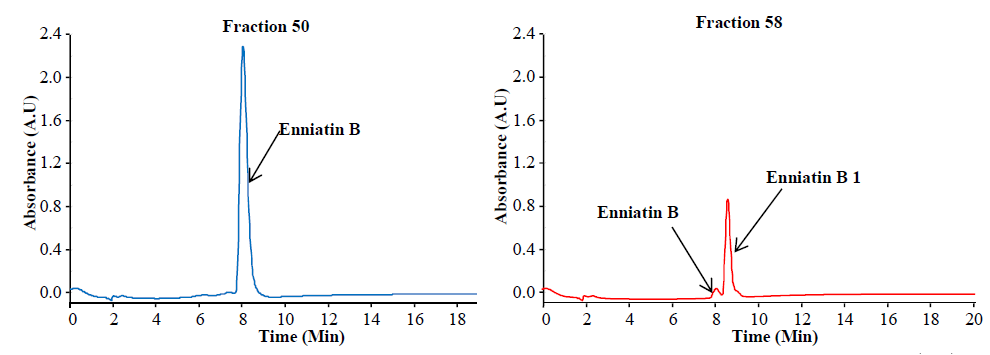
Preparative separation of individual enniatins from Fusarium tricinctum (Dr. Rosine Suchfort)
Flash chromatography is the fundamental method for natural product purification. We usually start with normal phase eluted with a gradient of cyclohexane, ethyl acetate and methanol.

Flash chromatography: in a fume hood you see the pumps (up to 5 MPa), the fraction collector and lower part of a self-filled column. The signal of the detector is displayed on a laptop screen.
Preparative HPLC

Our HPLC is from Jasco. You see the pumps and detector (left), a 2.5 cm preparative column (middle) and the fraction collector (right).
Preparative TLC is a simple yet very useful method, especially when many samples have to be screened fast. Preparative TLC is often used in our lab, too:

TLC box in a foome hood, with Dr. Yi Kuang preparing samples for loading
Analysis of volatiles compounds by GC
Some plant and fungal metabolites are volatile. These compounds are sampled from the gaseous phase and analyzed by GC. Directly in the lab we use a GC-FAIMS system which allows us to distinguish among isomers. The choice of software for data processing in GC-FAIMS is limited, we therefore use Perl to process raw data and generate chromatograms, as shown below on data collected by Johannes Ott for fungal VOCs:
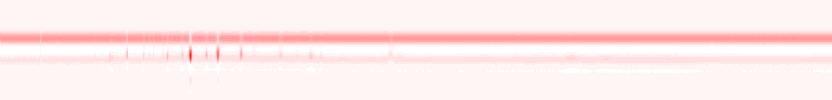

Raw histogram of GC-FAIMS with positive ionization (Johannes Ott)

Raw histogram of GC-FAIMS with negative ionization (Johannes Ott)
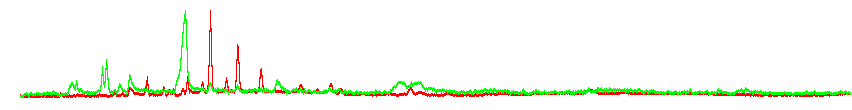
Normalized GC-FAIM chromatograms for positive (red) and negative (green) ionization; notice signals in neg. ionization hardly visible in raw histogram (Johannes Ott and Petr Karlovsky)
In most projects on VOCs we rely on the GC-MS equipment in the laboratory of Prof. Ivo Feußner. Thanks to the suspension bridge connecting our labs, moving forth and back between the labs is comfortable:

Bridge connecting our lab (left) with the lab of Prof. Ivo Feußner (right), photo by Petr
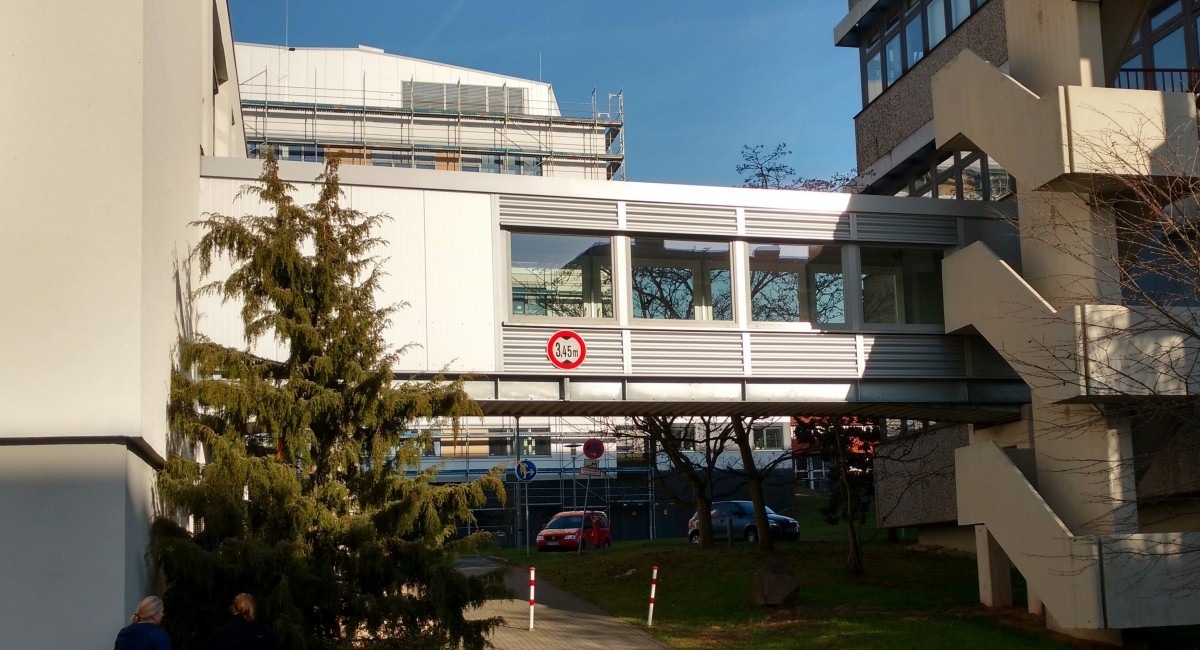

Bridge from the other side, photo by Franz
Georg-August-University Goettingen
Grisebachstrasse 6, 37077 Goettingen, Germany
FAX: +49 551 398177